Abstract
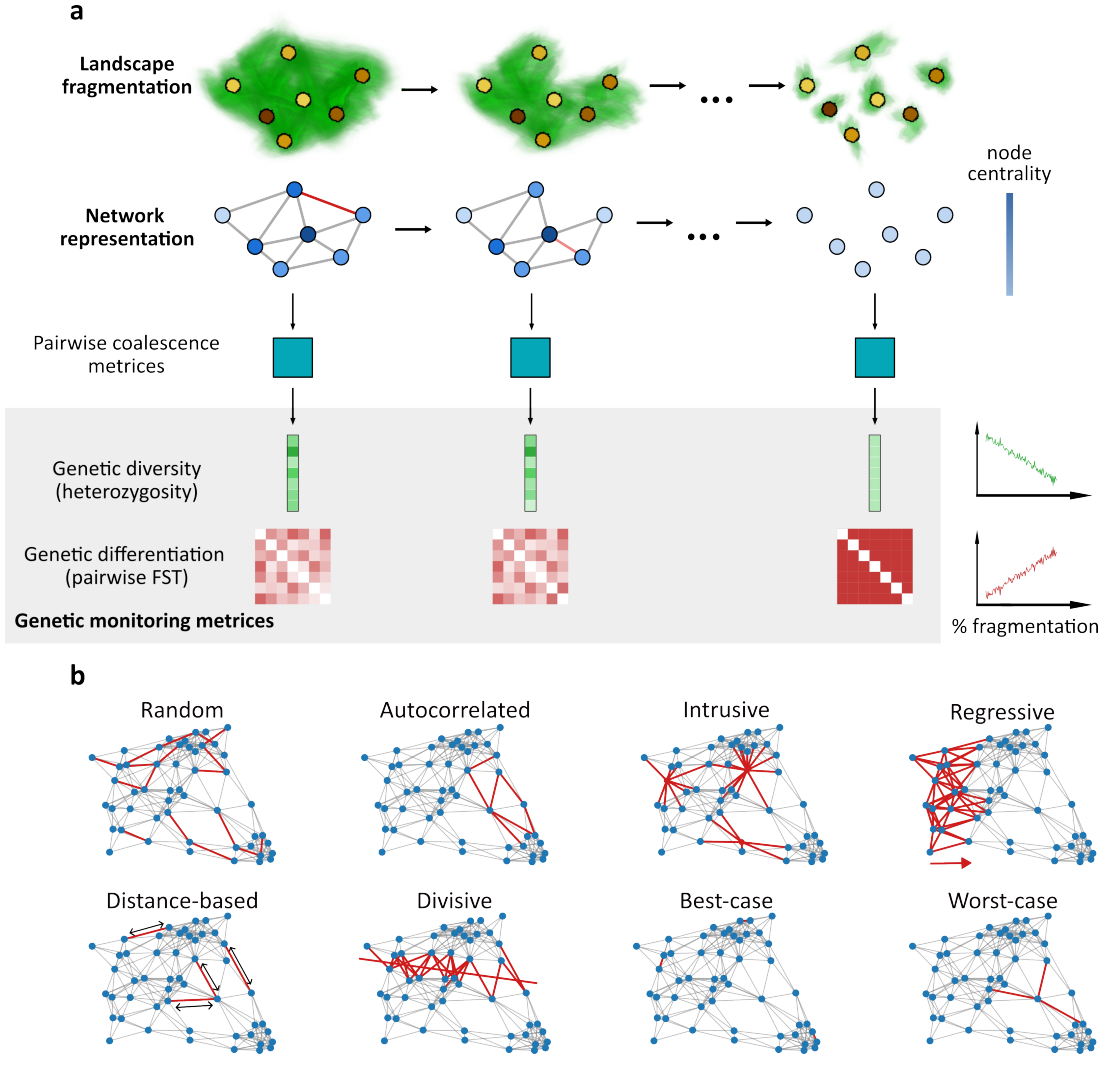
Habitat fragmentation is one of the most immediate and substantial threats to biodiversity, generating isolated populations with reduced genetic diversity. Genetic monitoring has become essential for detecting fragmentation and tracking its progress. However, the coherentinterpretation of geneticmonitoring data and understanding the genetic consequences of fragmentation require frameworks that accurately represent real-world complexity. Existing theoretical frameworks typically rely on simplified spatial structures and do not adequately capture the heterogeneous migration patterns of natural populations. Here,we integrate network theory andmathematical population geneticsto develop a framework forstudying the genetic consequences of fragmentation processes, explicitly accounting for heterogeneous connectivity and temporal dynamics. We apply this framework to examine how different fragmentation processes affect genetic measures commonly used in genetic monitoring. Through analysis of simulated and empirical networks, we find that different fragmentation scenarios produce substantially distinct trajectories in key genetic measures, sometimes exhibiting rapid transitional dynamics. Furthermore, fragmentation can cause deviations from classical theoretical expectations, such as the expected correlation between genetic and geographic distance (isolation-by-distance) or between genetic diversity and connectivity. Finally, we propose and demonstrate detectable early warning signals in genetic monitoring data that precede rapid transitional phases. Our framework thus provides a practical interpretation of genetic monitoring data, and a proof-of-concept that bridges the gap between idealized theoretical models and real-world connectivity dynamics.
Press Coverage
- A New Tool Predicts Wildlife “Genetic Collapse” Before It’s Too Late, A–Z Animals (25 March 2026).